SAMRC Wastewater Surveillance and Research Programme
Frequently Asked Questions (FAQs)
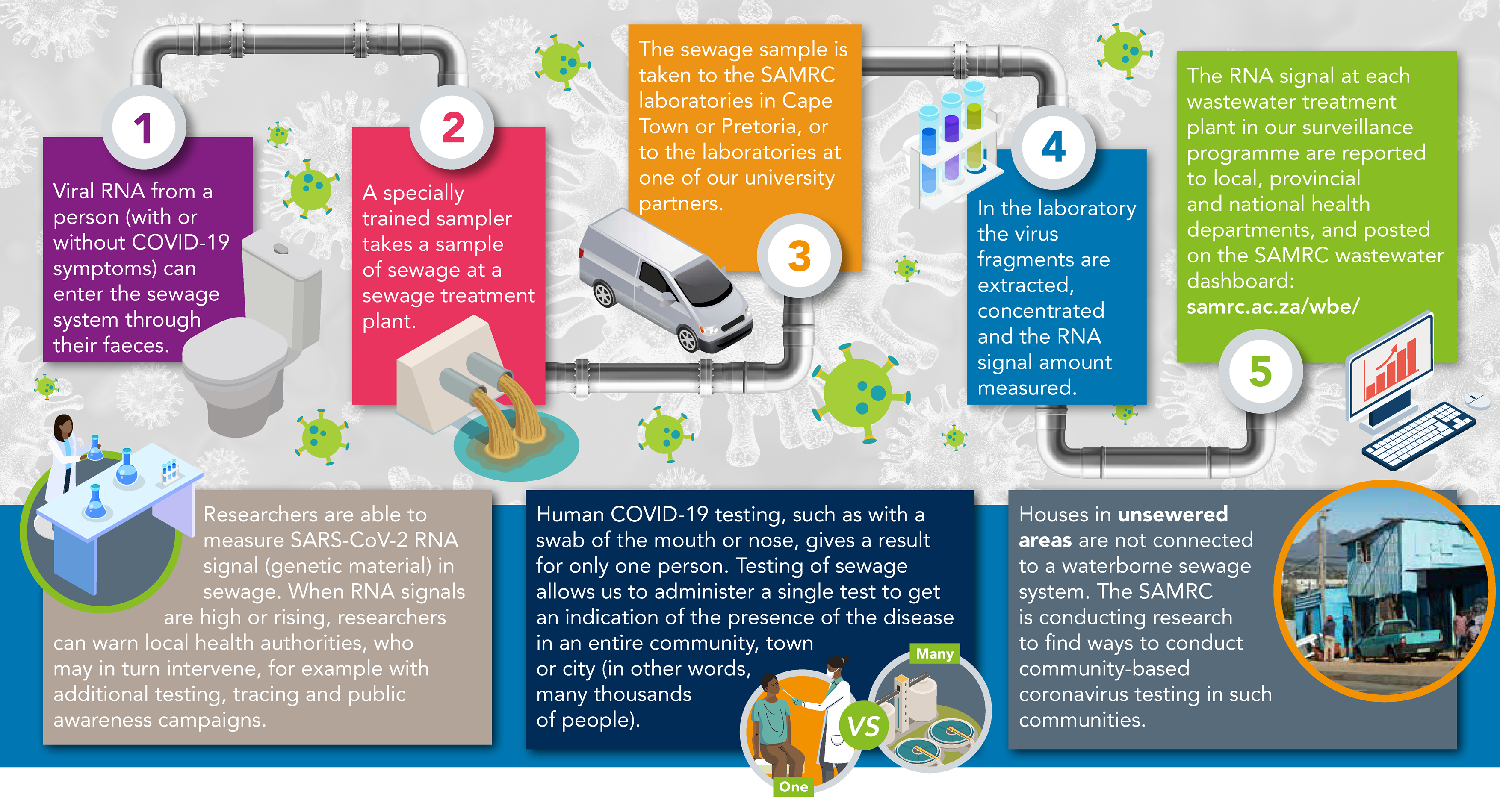
On Monday mornings, a single (grab) wastewater sample is collected from selected wastewater treatment works (WWTWs) in various parts of the country by trained wastewater samplers. As soon as possible after collection, the samples are transported under refrigerated conditions to the SAMRC or partner university-based laboratories. Please refer to Wastewater Sampling Guide that has been developed for the purposes of the SAMRC Wastewater Surveillance and Research Programme (SARS-CoV-2 Wastewater Sampling Guide)
The samples are analysed at the SAMRC or partner university-based laboratories. To date, published research indicates that the virus is inactivated (no longer infectious) when the RNA fragments are detected in the wastewater. The laboratory confirms the presence (qualitative analysis) and determines the SARS-CoV-2 RNA signal by reverse transcription polymerase chain reaction (RT-PCR) (quantitative) analysis. Results from two gene probes are aggregated and presented as the RNA signal copies per millilitres (copies/ml) found in wastewater sample.
The results are provided to our public health stakeholders across the country within 48 hours of sample collection and are also shared on our public dashboard. The SARS-CoV-2 RNA signal is presented in categories (Category 1 = lowest SARS-CoV-2 RNA signal, Category 5 = highest SARS-CoV-2 RNA signal). The trends observed in the wastewater may give us an indication of COVID-19 case trends in a particular wastewater catchment area. This becomes very useful to identify COVID-19 hotspots and can guide action in preparedness strategies for COVID-19.
There are several limitations to wastewater surveillance in terms of COVID-19. In the course of a day or week, people may travel across several wastewater catchment areas for the purposes of work, education, recreation, seeking health care, shopping, visiting friends and family and many other reasons. In this way they may contribute to multiple wastewater catchment areas. Another factor is uncertainties in shedding of the virus; the amount and duration of virus shedding is still unclear. Therefore, currently, it is not possible to accurately predict the number of infected individuals within a particular wastewater catchment area or community, based on wastewater surveillance.
Participating SAMRC Units and Platforms
.png) |
.png) |
 |
 |
||||
| Environment & Health Research Unit | Biomedical Research and Innovation Platform | Biostatistics Research Unit | Genomics Centre |




